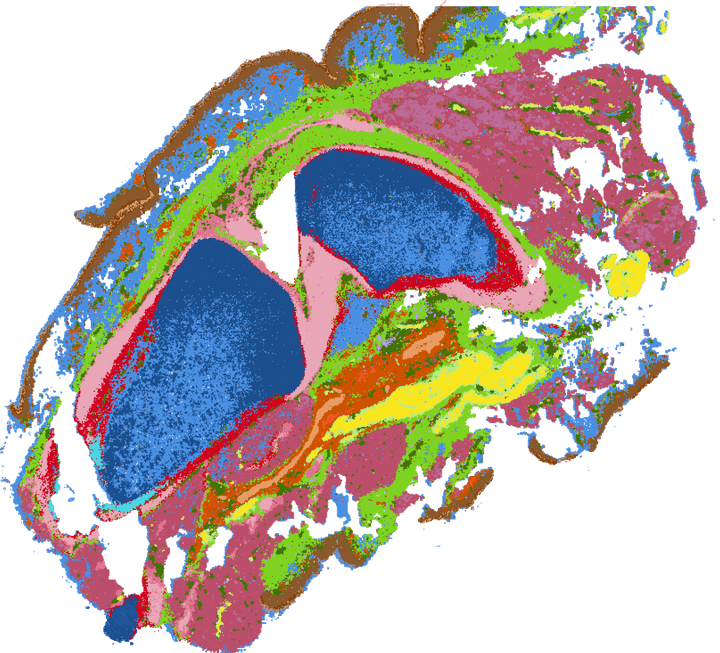
Spatial transcriptomics of developing human knee
I study how novelties arise over the course of evolution and development — using the vertebrate skeleton as a window into 400 million years of change.
Research Focus
Evolutionary noveltiesSpecies across deep time
- Homo sapiens — Human, extant
- Xenopus tropicalis — Frog, 350 mya
- Danio rerio — Zebrafish, 420 mya
- Petromyzon marinus — Lamprey, 500 mya
Anatomical focus in Homo sapiens
- Pelvis — Re-architected for bipedalism — the load-bearing hinge between spine and legs.
- Hip — Ball-and-socket joint where the femoral head meets the acetabulum — remodelled for upright gait and weight bearing.
- Hyoid — Throat skeleton derived from ancestral gill arches. Central to speech and swallowing.
- Knee — The largest synovial joint — cartilage, menisci, and ligaments under constant load.
Methods applied across all organisms
- DNA / Genetics — Comparative single-cell RNA sequencing integrated with spatial transcriptomics to study the underlying gene expression patterns.
- CRISPR — Targeted gene editing to test developmental hypotheses.
- CT scanning — Non-destructive 3D imaging of skeletal morphology in embryos and comparative museum specimens.
- Histology — Tissue-level architecture of bone, cartilage, and ligament using staining methods like trichrome and H&E.
- ATAC-seq — Open-chromatin profiling at single-cell level to study which regulatory regions are active during skeletal morphogenesis.
- Single-cell — Generate cell-type atlases across development and species to understand deep homology.
Evolution has produced remarkable structures — from the fused tail rod that lets frogs jump to the broad pelvis that lets humans walk upright. These are evolutionary novelties: structures that appear in one group of animals but have no clear counterpart in their ancestors.
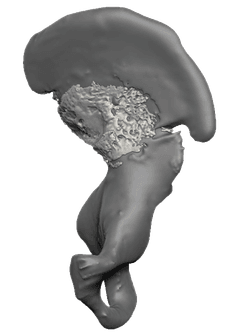
Evolution of human bipedalism
A multifaceted study — histology, comparative genomics, and spatial transcriptomics across 100+ embryonic tissue samples from humans and ~24 primate species — uncovering the developmental basis of the uniquely human pelvis.
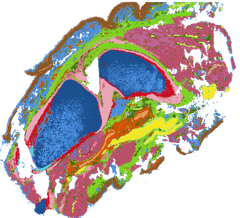
Evo-devo of bipedal joints
Two clinically critical synovial joints — hip and knee — examined through scRNA-seq, scATAC-seq, and high-resolution spatial transcriptomics to map joint-specific cellular and regulatory atlases.
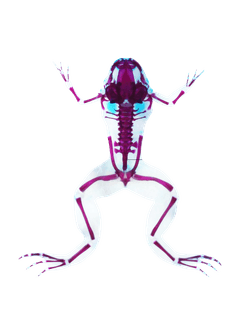

Developmental origins of the anuran urostyle
The urostyle — a fused bony rod at the end of the frog vertebral column — is an evolutionary novelty that arose nearly 200 million years ago. Its developmental origins had long been poorly understood.