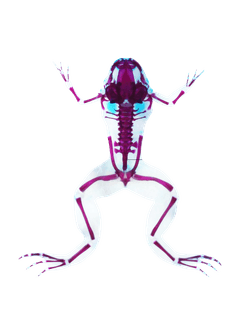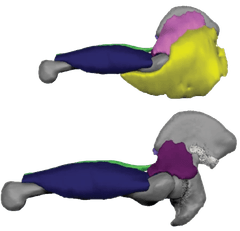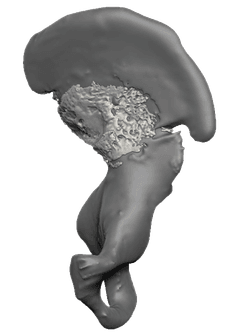
Evolution of human bipedalismand the pelvic girdle
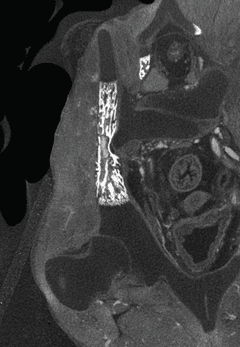
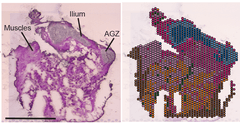
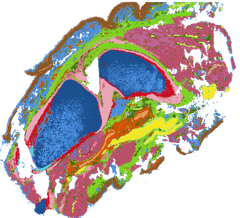
A multifaceted study — histology, comparative genomics, and spatial transcriptomics across 100+ embryonic tissue samples from humans and ~24 primate species — uncovering the developmental basis of the uniquely human pelvis.
- Heterotopy
- A 90° rotation of the cartilage growth plate produced a wide ilium instead of the tall, narrow blade seen in other primates.
- Heterochrony
- A shift in the timing and location of bone formation (ossification) reorganised how the ilium ossifies during development.
- Genes
- 300+ genes drive these changes. SOX9 and PTH1R control growth-plate reorientation; RUNX2 regulates the ossification shift.
- When
- The shifts likely began 5–8 million years ago, around our split from the African apes — enabling bipedal locomotion.